- 4192 Devonwood Way, Ashburn, Virginia, 20148
- Helpline: +1 (703) 665-3747
Immunogenetic Studies and Humoral Reactivity in an Affected COVID-19 First Wave Family
- Home
- Back to Journal
- Article Details
Introduction
The Covid-19 pandemic has been divided into different infection periods (waves) [1]. The first wave started in Europe mid-January 2020 and was characterized initially by dedicated hotspots of infections, from which the virus spread to the general population [2]. The second and the possibly third wave are characterized by viral mutations and possible increased virulence. Even during the first wave mutations like the spike gene D614G with higher infectiousness evolved [3]. Earlier it was stated that blood group O might be to a certain degree protective against SARS-CoV-2 infection or at least protects from a severe course of COVID-19 [4]. We started a local cooperative study on the influence of immunogenetic factors on disease occurrence, spreading, and severity. Here, we report on HLA, blood group, and humoral antiviral immunity data of a SARS-CoV-2-infected family during the first wave of the pandemic disease.
Material and Methods
Our laboratory is accredited for the PCR technology and antibody detection by Luminex technology by the European Federation for Immunogenetics and participated successfully in several external proficiency testing exercises on HLA typing by Next Generation Sequencing (NGS). Regarding this technology we used the AlloSeq® Tx 17 from CareDx, San Fransisco, CA, USA, for HLA typing. The sequencing was performed on Miseq from Illumina (San Diego, CA, USA), following strictly the manufactures recommendations. Alternatively Real Time PCR was used. In addition to the “classical” loci HLA-A, B, C, DRB1, DRB3, DRB4, DRB5 DQA1, DQB1, DPA1 and DPB1, also typed for HLA-E, HLA-F, HLA-G, HLA-H, MICA and MICB. The results of HLA-F and H were not informative and therefore not reported here.
ABO and Rh typing were performed using the PCR sequence specific priming for blood groups of BAG, Lich, Germany, following the manufactures recommendations.
The LAB ScreenTM COVID Plus assay is Luminex® based from One Lambda, West Hills CA, USA. It was strictly followed the manufactures recommendations and used the proposed cutoff values as given by the manufacturer to quote a given value as positive or negative. The assay consists of antigenic determinants bound to polystyrene beads and used in the Luminex® set up (LX200, IS 2.3). The assay detects simultaneously the presence of 5 different SARS-CoV-2 targets and 6 additional coronavirus targets for antibodies present in the subject serum (modified from One Lambda (https://www.onelamdba.com)).
The study was accepted by the ethical committee of our hospital (195/20e-ek). After detailed explanation of the study and investigation the agreement of the family´s legal guardians was obtained by signing the informed consent. For the study, EDTA and serum samples were taken according to the study documents (provided as supplemental material) in case of need.
Results
Family LEI-01:
The family consists of the mother (M), her two daughters (C1, 16 years old and C2, 13 years old) from a previous partnership, her current partner (F2 and a mutual daughter (F2M C1, 17 months). All members of the family were tested by SARS-CoV2 PCR assay according to WHO standards (https://www.WHO.int; https://www.ecdc.europa.eu/en/covid-19/latest-evidence/diagnostic-testing). All were positive with exception of C1. All family members were admitted to the hospital. F2 had severe respiratory problems with fever and oxygen was administrated. M had mild cold symptoms with pneumonia but no fever. C2 had mild fever and cold symptoms, while C1 and F2M C1 were asymptomatic (Table 2). NB the family shares housing. HLA and blood group typing were performed by molecular means. In addition, a novel Covid antibody assay was used, in which five SARS-CoV2 specific antigens was included as targets for the antibodies, and six additional human specific antigens of other coronavirus than SARS-CoV2 were used in parallel (Table 1).
Table 1: Antibody targets in the assay.
|
Target |
Description |
|
SARS-CoV-2 Spike |
Extra Cellular Domain (ECD) |
|
SARS-CoV-2 SpikeS1 |
S1 Domain |
|
SARS-CoV-2 Spike RBD |
Receptor Binding Domain |
|
SARS-CoV-2 SpikeS2 |
S2 Domain |
|
SARS-CoV-2 Nucleocapsid Protein |
Nucleocapsid Protein |
|
HCOV-229E Spike S1 |
Common Cold Human Coronavirus 229E |
|
HCoV-HKUl SpikeS1 |
Common Cold Human Coronavirus Hong Kong Ul |
|
HCOV-NL63 Spike S1 |
Common Cold Human Coronavirus Netherland 63 |
|
HCOV-OC43 Spike S1 |
Common Cold Human Coronavirus Organ Culture43 |
|
MERS-CoV Spike S1 |
Middle East Respiratory Syndrome Coronavirus |
|
SARS-CoV Spike S1 |
Severe Acute Respiratory Syndroma Coronavirus (2003) |
Table 2: Symptoms of family members who have Covid-19.
|
|
F2 |
M |
F2M C1 |
C2 |
|
duration of symptoms |
10 days |
14 days |
no* |
1 day |
|
fever |
yes, max. 39,5°C |
no |
no* |
yes, 38°C |
|
headache |
yes |
yes |
no* |
not known |
|
body aches |
yes |
yes |
no* |
not known |
|
tiredness |
yes |
yes |
no* |
not known |
|
cough |
yes |
no |
no* |
not known |
|
runny nose |
no |
yes |
no* |
not known |
|
hoarseness |
no |
no |
no* |
not known |
|
sore throat |
no |
no |
no* |
not known |
|
pneumonia |
yes |
yes |
no* |
not known |
|
sinus discomfort |
no |
no |
no* |
not known |
|
dizziness |
no |
no |
no* |
not known |
|
taste/smell disorders |
yes, since day 4 until January 2021 |
yes |
no* |
yes, lightly |
|
depressive moods |
no |
no |
no* |
not known |
|
skin redness |
no |
no |
no* |
not known |
|
skin vesicles |
no |
no |
no* |
not known |
|
nausea/vomiting |
no |
no |
no* |
not known |
|
abdominal pain |
no |
no |
no* |
not known |
|
diarrhea |
yes |
no |
no* |
not known |
|
constipation |
no |
no |
no* |
not known |
|
hair loss |
yes |
yes |
no* |
not known |
|
Hospitalized, hospital days |
yes/16 days |
yes/not known** |
yes/not known** |
not known |
|
oxygen supply |
yes, 2l/min |
no |
no* |
not known |
|
Invasive/non-invasive ventilation |
no |
no |
no* |
not known |
|
medication for treatment during infection |
not known |
not known |
none |
not known |
|
long-term medication |
none |
Cardesartan |
none |
not known |
|
other |
fungal infection |
hip pain, increased blood pressure, myocarditis |
no* |
not known |
*) observations of the parents; **no treatment needed
The results of the HLA typing of the family are depicted in Table 3. The HLA haplotypes of the first partner of the mother are deduced. The F2M C1 child was typed for HLA-A, B, C, DRB1 and DQB1 in 1-field resolution. The remaining results were deduced from the inherited haplotype C from M and E from F2. No sharing of haplotypes was observed. However, since the C1 and C2 daughters inherited different haplotypes from the mother one could speculate that an immunogenetic component might influence the severity of the Covid-19 infection. The asymptomatic F2M C1 child inherited the same haplotype as the C1 and could represent a protective haplotype.
Table 3: HLA typing of the family LEI-01.
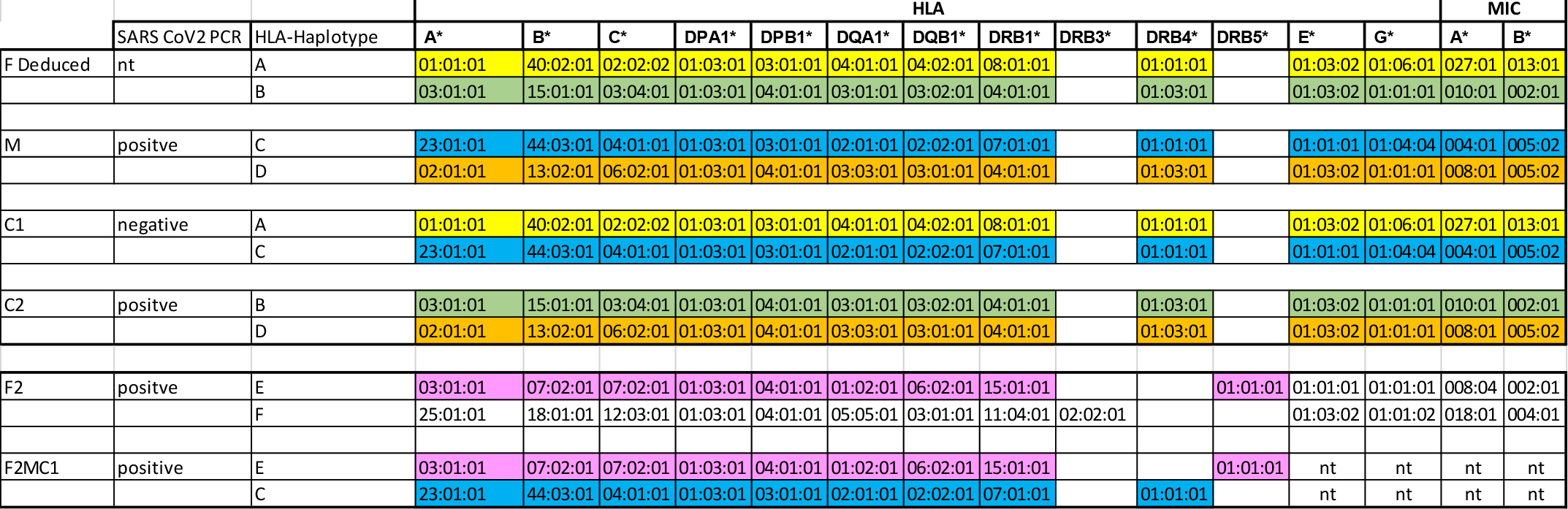
We also performed the definition of the blood groups by sequence specific priming PCR, with exception of F2 and M. Their typing was done by usual serological manner. The results are shown in Table 4. Blood group O was shown to be rather protective towards SARS-CoV2 [4]. Since all members of the family are BGO no conclusions are drawn.
Table 4: ABO and Rh typing of the family.
|
|
SARS CoV2 PCR |
ABO |
|
RH |
|
F2 |
Positive |
0 |
nd |
CcDEe |
|
M |
Positive |
0 |
nd |
CCDee |
|
F2M Cl |
Positive |
01 |
01v |
CcDDEe |
|
Cl |
Negative |
01 |
01v |
CCDDee |
|
C2 |
Positive |
01 |
01v |
CCDDee |
Finally, the reactivity of the serum antibodies towards the different SARS-CoV-2 antigens and those from epidemic coronaviruses were analyzed. The results are shown in Table 5.
Table 5: Antibody reactivity towards SARS-CoV2 and other coronavirus antigens (shaded boxes indicate positive results).
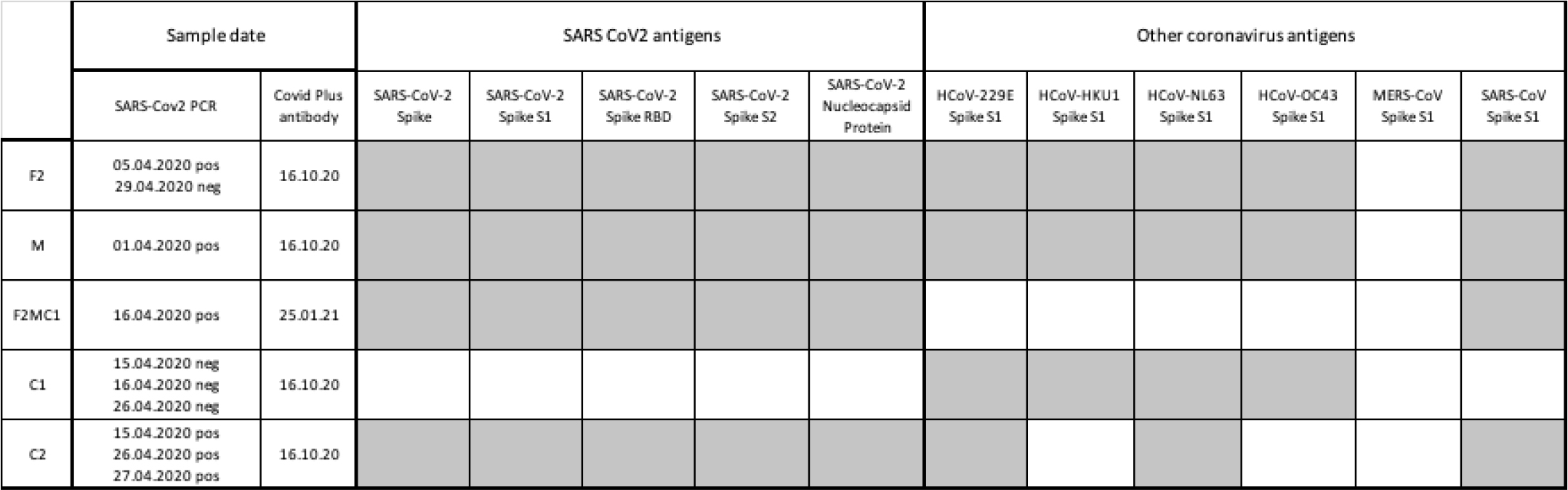
All members of the family with exception of the C1 showed positive reactions against all five SARS-CoV2 targets. Because of the negative PCR assay, a negative antibody assay for C1 is plausible.
Discussion
The family LEI-01 came to our Institution to participate in our local cooperative study. The members of this family live together and are prone to cluster infections like COVID-19. The C1 individual had no infection as proven by negative PCR and lack of antibodies against the SARS-CoV-2 while antibodies against epidemic coronaviruses were detected. M and the F2M C1 child had mild symptoms or no symptoms at all. The mother was PCR positive and developed antibodies against several SARS-CoV-2 antigens, as also her F2M C1 offspring. The latter had no symptoms.
Very little is known with regard to SARS-CoV-2 infection and the Major Histocompatibility Complex [5–7]. It is far too early to define immunogenetic factors in the family presented where the respective individuals had varying degrees of symptoms. However, because of the different affected HLA haplotypes one could speculate that the haplotype inherited from the mother to the C1 child might include a protective factor. These three individuals express the haplotype C including the HLA-A*23:01, B*44:03, C*04:01, DRB1*07:01, DQA1*02:01, DQB1*02:02. We can speculate that this haplotype protects in a way the carriers from SARS-CoV2 infections and severity. Further analyses in a wider patient population will show whether this assumption is correct.
Acknowledgement
We would like to thank the family and the family members for their acceptance to participate in the study.
References
- Cacciapaglia G, Cot C, Sannino F (2020) Second wave COVID-19 pandemics in Europe: a temporal playbook. Sci Rep 10: 15514. [View]
- Robert Schlack, Laura Neuperdt, Heike Hölling, et al. (2020) Impact of the COVID-19 pandemic and the related containment measures on the mental health of children and adolescents. Journal of Health Monitoring · 2020 5(4) , RKI, Berlin. [View]
- Plante JA, Liu Y, Liu J, et al. (2020) Spike mutation D614G alters SARS-CoV-2 fitness. Nature. [View]
- David Ellinghaus, Frauke Degenhardt, Luis Bujanda, et al. (2020) Genomewide Association Study of Severe Covid-19 with Respiratory Failure. N Engl J Med. 383: 1522–1534. [View]
- Petra Bacher, Elisa Rosati, Daniela Esser et al. (2020) Low-Avidity CD4+ T Cell Responses to SARS-CoV-2 in Unexposed Individuals and Humans with Severe COVID-19. Immunity 53: 1258–1271.e5. [View]
- Alba Grifoni, Daniela Weiskopf, Sydney Ramirez, et al. (2020) Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals. Cell 181: 1489–1501.e15. [View]
- Salman Toor, Reem Saleh, Varun Sasidharan Nair, et al. (2020) T?cell responses and therapies against SARS?CoV?2 infection. Immunology 162: 30–43. [View]
Article Type
Case Report
Publication history
Received: February 08, 2022
Accepted: February 26, 2022
Published: March 08, 2022
Citation:
Lehmann Claudia, Landgraf Ramona, Lakowa Nicole, Grünewald Thomas, Henschler Reinhard and Doxiadis Ilias (2022) Immunogenetic Studies and Humoral Reactivity in an Affected COVID-19 First Wave Family. Med Case Rep Ther Stud 03(01): 67–71.
Lehmann Claudia1, Landgraf Ramona1, Lakowa Nicole2, Grünewald Thomas2, Henschler Reinhard1 and Doxiadis Ilias1
1Laboratory for Transplantation Immunology, Institute of Transfusion Medicine, University Hospital Leipzig, Germany
2Clinic for Infectious Diseases and Tropical Medicine, Klinikum Chemnitz, Germany
*Corresponding author
Lehmann Claudia,
Laboratory for Transplantation Immunology,
Institute of Transfusion Medicine,
University Hospital Leipzig,
Institute of Transfusion Medicine,
Johannisallee 32,
04103 Leipzig,
Germany;
Tel: +49 341 9725350;